Emergent Biological Realism in RL-Trained DNA Language Models
Emergent Biological Realism in RL-Trained DNA Language Models
Thiel, M.; Cunningham, A.; Barnes, C. P.
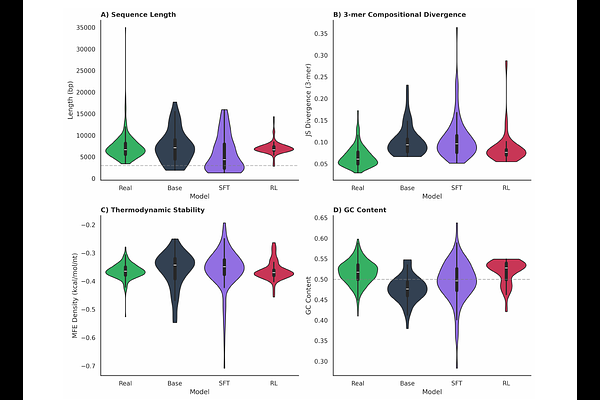
AbstractReinforcement learning has driven the mass adoption of large language models by unlocking unexpected capabilities, yet this approach remains largely underexplored for generative DNA models. We investigate whether similar post-training techniques can induce emergent biological realism in DNA language models, using plasmid generation as a testbed due to plasmids' relative simplicity, well-characterized functional constraints, and ubiquity in biotechnology. Using Group Relative Policy Optimization with a reward function based on constraints from engineered biology, our model achieves a 77% quality control pass rate compared to 5% for the pretrained baseline. Remarkably, beyond explicitly optimized features, the model exhibits surprising biological parallels: generated sequences match natural plasmids in thermodynamic stability, codon usage patterns, and ORF length distributions, properties not explicitly optimized in the reward function. These results suggest that RL post-training can steer DNA language models toward biologically coherent regions of sequence space, analogous to how such techniques unlock unexpected capabilities in natural language models, particularly in verifiable domains.