MEIsensor: a deep-learning method for mobile element insertion discovery
MEIsensor: a deep-learning method for mobile element insertion discovery
Wang, Y.; Zhang, P.; Wan, S.; Zhang, Z.; Sun, P.; Xu, T.; Jia, P.; Ye, K.; Yang, X.
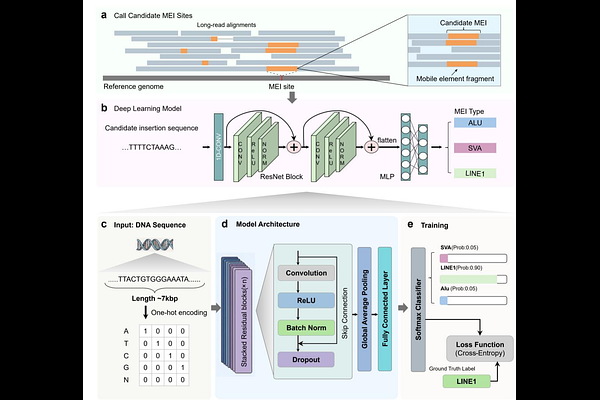
AbstractMobile element insertions (MEIs) are a critical source of structural variation in the human genome, yet their accurate detection remains challenging, particularly within repetitive regions and for structurally complex insertions. While long-read sequencing enables the direct recovery of inserted sequences, current analytical pipelines rely heavily on repeat-library alignment, which can limit both computational efficiency and accuracy. Here, we present MEIsensor, a deep learning framework designed to detect and classify MEIs directly from long-read sequencing data. Unlike conventional methodologies, MEIsensor performs direct, sequence-based annotation of Alu, LINE1, and SVA insertions. Evaluated against HGSVC benchmarks using HiFi long-read datasets, MEIsensor consistently outperformed existing tools across major MEI classes while substantially improving computational efficiency, demonstrating particularly pronounced gains for structurally complex SVA insertions. Furthermore, manual curation confirmed that MEIsensor successfully identifies MEIs hidden in highly repetitive genomic regions, including centromeric higher-order repeats, that were previously absent from benchmark datasets. Together, these advancements establish MEIsensor as an efficient framework for MEI analysis that promises to significantly advance our understanding of human genomic architecture and evolution.