Exploring Chromosomal Position Effects for Predictable Tuning of Metabolic Pathways in Yeast
Exploring Chromosomal Position Effects for Predictable Tuning of Metabolic Pathways in Yeast
Hong, H.; Cai, Y.; Wang, J.; Dong, C.; Zhang, Q.; Lian, J.
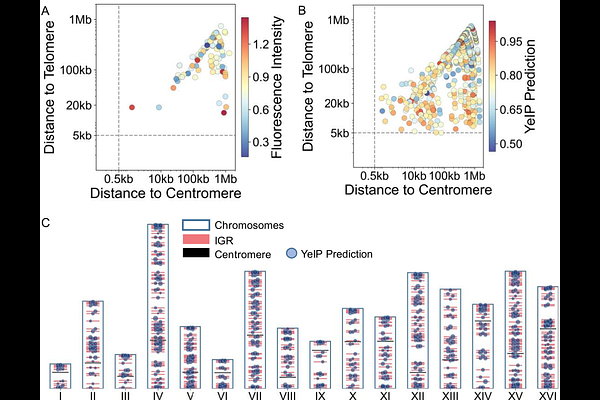
AbstractPrecise control of gene expression is essential for optimizing metabolic pathways, yet current tuning strategies based on promoter strength or gene copy number remain largely empirical. Chromosomal position represents an additional regulatory axis, as identical gene expression cassettes can exhibit markedly different expression levels depending on their integration sites. Here, we systematically quantified the expression output of 98 intergenic regions (IGRs) in Saccharomyces cerevisiae using a fluorescent reporter and developed a predictive framework, Yeast IGR Prophet (YeIP), that infers expression potential directly from genomic context. By integrating multi-scale genomic features including transcriptional neighborhood, chromatin state, and chromosome topology, YeIP accurately predicted expression ranks and enabled the construction of a genome-wide atlas of expression hotspot and coldspot. Using this atlas, we rationally optimized a three-gene lycopene pathway solely through genomic integration site selection, achieving optimal transcriptional stoichiometry without modifying promoters or gene copy numbers. These results transform chromosomal integration sites from static genomic coordinates into programmable regulatory elements, establishing a predictive, data-driven framework for rational and scalable design of metabolic pathways in yeast.