Nextstrain automates real-time phylodynamic analysis of open data for endemic and emerging pathogens
Nextstrain automates real-time phylodynamic analysis of open data for endemic and emerging pathogens
Andrews, K. R.; Chang, J.; Roemer, C.; Hadfield, J.; Lin, V.; Brito, A. F.; Daodu, R.; Joia, I. A.; Kistler, K.; Li, A. W.; Moncla, L. H.; Paredes, M. I.; Kuhnert, D.; Torres, L. M.; Voitl, L.; Aksamentov, I.; Hodcroft, E. B.; Huddleston, J.; McCrone, J. T.; Anderson, J. S.; Sibley, T. R.; Lee, J.; Neher, R. A.; Bedford, T.
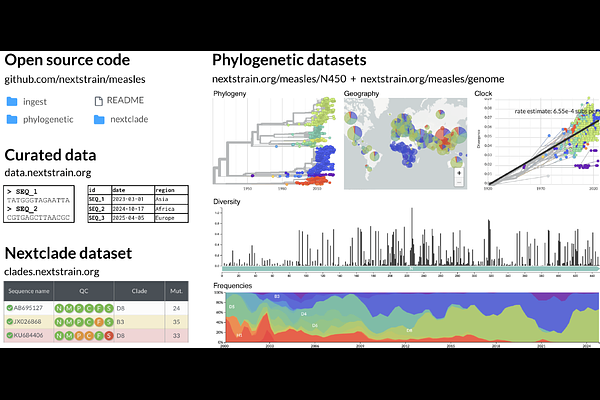
AbstractMotivation: Genome sequencing provides an exceptional window into the evolutionary and epidemiological dynamics of endemic and emerging pathogens, and thus allows for better, more targeted, public health interventions. Online genomic surveillance platforms can provide near real-time insight into these dynamics. Results: Nextstrain provides continually updated real-time genomic surveillance for 21 viruses and the bacterial pathogen Mycobacterium tuberculosis, with most analyses relying solely on open sequence data. Each pathogen includes steps to fetch and curate open data, classify sequences using established nomenclature systems, perform phylodynamic analyses, and share the results publicly. These analyses are automated, with most running daily to provide continually updated snapshots of pathogen evolution. Availability and Implementation: All source code is available at https://github.com/nextstrain. Phylodynamic results can be visualized and downloaded at https://nextstrain.org/pathogens, and open sequence data and curated metadata are available at https://nextstrain.org/pathogens/files.