CPS: Mapping Physical Coordinates to High-Fidelity Spatial Transcriptomics via Privileged Multi-Scale Context Distillation
CPS: Mapping Physical Coordinates to High-Fidelity Spatial Transcriptomics via Privileged Multi-Scale Context Distillation
Zhang, L.; Cao, K.; Zheng, S.; Liang, S.; Wan, L.
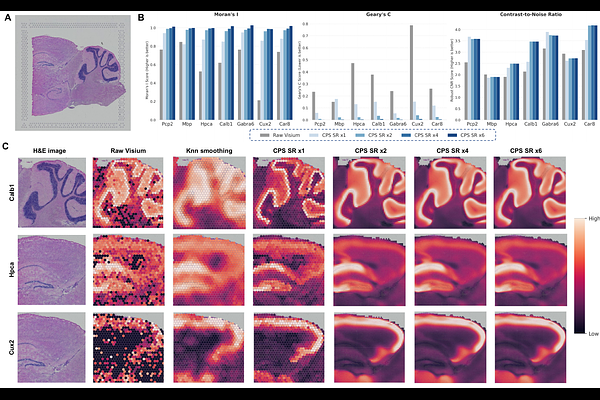
AbstractMotivation: Spatial transcriptomics enables the dissection of tissue heterogeneity within native contexts, yet current platforms are inherently constrained by high sparsity and low signal-to-noise ratios that obscure fine-grained biological signals. Current efforts to recover these signals are limited by image registration dependencies or the inherent context-blindness of implicit neural representations. Results: We introduce the Cell Positioning System (CPS), a context-aware implicit neural representation framework designed to map physical coordinates to high-fidelity spatial transcriptomics via a privileged multi-scale context distillation strategy. CPS treats multi-scale tissue niches as privileged information, employing a teacher network equipped with a multi-scale niche attention mechanism to capture adaptive biological interactions during training. This structural knowledge is explicitly distilled into a student coordinate network, enabling the generation of context-aware expression landscapes solely from spatial coordinates during inference. Benchmarking on the DLPFC dataset demonstrates that CPS achieves state-of-the-art performance in spatial and gene expression imputation and denoising. Furthermore, CPS enables super-resolution to recover high-resolution mouse brain anatomical details and offers interpretability by identifying the scale effective size of biological interactions within human breast cancer tissues. Finally, the framework exhibits superior scalability for large-scale datasets with linear computational complexity.