BIOFORMERS: A SCALABLE FRAMEWORK FOR EXPLORING BIOSTATES USING TRANSFORMERS
BIOFORMERS: A SCALABLE FRAMEWORK FOR EXPLORING BIOSTATES USING TRANSFORMERS
Belgadi, S. A.; Li, O.; Zhang, D. Y.; Gopinath, A.
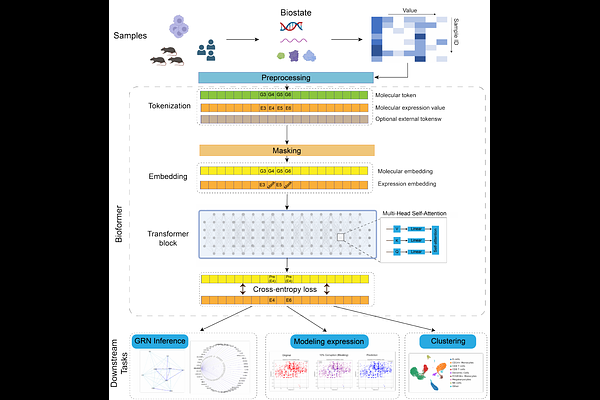
AbstractGenerative pre-trained models, such as BERT and GPT, have demonstrated remarkable success in natural language processing and computer vision. Leveraging the combination of large-scale, diverse datasets, transformers, and unsupervised learning, these models have emerged as a promising method for understanding complex systems like language. Despite the apparent differences, human language and biological systems share numerous parallels. Biology, like language, is a dynamic, interconnected network where biomolecules interact to create living entities akin to words forming coherent narratives. Inspired by this analogy, we explored the potential of using transformer-based unsupervised model development for analyzing biological systems and proposed a framework that can ingest vast amounts of biological data to create a foundational model of biology using BERT or GPT. This framework focuses on the concept of a `biostate,\' defined as a high-dimensional vector encompassing various biological markers such as genomic, proteomic, transcriptomic, physiological, and phenotypical data. We applied this technique to a small dataset of single-cell transcriptomics to demonstrate its ability to capture meaningful biological insights into genes and cells, even without any pre-training. Furthermore, the model can be readily used for gene network inference and genetic perturbation prediction.