PhagePickr: A bacteria-centric computational tool for designing evolution-proof phage cocktails
PhagePickr: A bacteria-centric computational tool for designing evolution-proof phage cocktails
Oneto, A.; Okamoto, K. W.
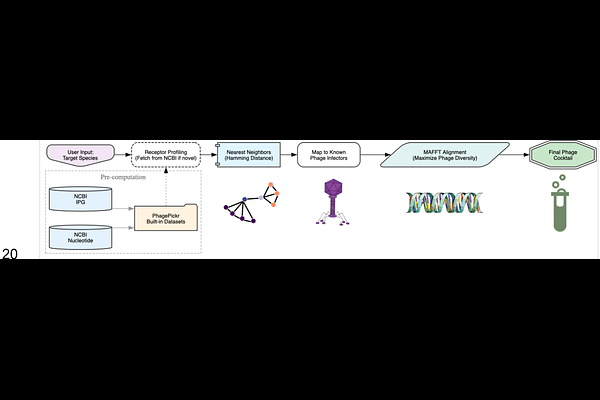
AbstractAs antibiotic resistance poses a major threat to global health, phage therapy offers an alternative to antibiotic treatments in the face of multidrug-resistant bacteria. However, host resistance to phages is also well-documented. Current computational tools for phage cocktail design do not explicitly address the evolution of phage resistance, let alone through the profiling of bacterial receptors whose variability drives much of phage resistance. We introduce PhagePickr, a computational pipeline for the automated design of phage cocktails that minimize host resistance. Unlike other tools, PhagePickr selects phages based on bacterial surface receptor similarity and prioritizes phage diversity to prevent cross-resistance. The tool uses NCBI datasets, a Nearest Neighbors algorithm, and Multiple Sequence Alignment to identify phenotypically similar hosts and ensure phylogenetic diversity in the final cocktail. We evaluated the utility of PhagePickr on ESKAPE pathogens and two understudied bacteria species. The cocktails included candidate phages predicted to target diverse receptors, comprising both lytic phages with confirmed therapeutic potential and novel candidates from similar species. We demonstrate the tools utility in generating cocktails and its capacity to scale as current databases are updated. PhagePickr provides a novel bacteria-centric framework for designing resistance-proof cocktails by exploring shared phenotypes.