Gene network centrality affects parallel evolution and local adaptation in wild yeast
Gene network centrality affects parallel evolution and local adaptation in wild yeast
Subramanian, S.; Bolnick, D. I.
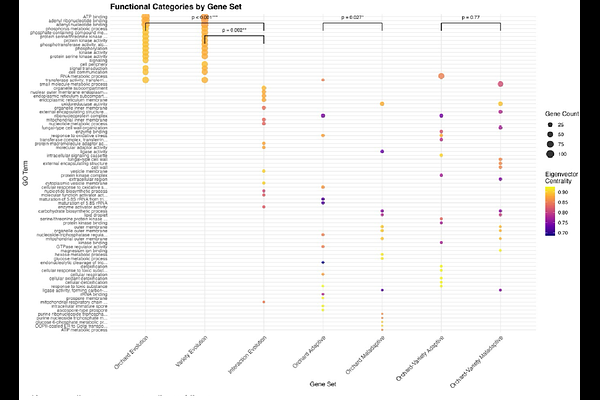
AbstractThe predictability of evolution remains a fundamental question in biology. While parallel evolution has been observed across taxa, we lack a mechanistic framework for predicting when evolution will be repeatable versus contingent. Here, we test whether gene network architecture predicts evolutionary repeatability and local (mal)adaptation in natural populations. We sequenced 38 wild Hanseniaspora uvarum yeast strains from two replicated apple varieties across four Connecticut orchards, yielding 29,489 high-quality SNPs. Population genomic analyses suggested that genetic differentiation was structured primarily by orchard environment. Reciprocal transplant experiments across all orchard-variety combinations demonstrated local (mal)adaptation at multiple ecological scales, with strongest (mal)adaptive responses at the orchard-variety level. By mapping H. uvarum genes to the comprehensive Saccharomyces cerevisiae genetic interaction network, we found that network centrality affects evolutionary outcomes. Highly connected genes occupying bow-tie network positions evolved in parallel across replicated apple varieties and central genes harbored alleles with adaptive fitness effects. In contrast, peripheral genes with fewer interactions facilitated rapid, non-parallel evolution to orchard-variety interactions and were associated with local maladaptation. Analysis of 150-gene sets evolving to apple variety within each orchard revealed greater-than-random similarity at genetic, functional, and network levels, with network-level predictability significantly exceeding gene-level predictability. These results suggest that gene network architecture could provide a mechanistic foundation for determining which evolutionary changes will be repeatable, potentially enabling evolutionary predictions even when specific genes cannot be identified.