Phyllosphere and rhizosphere microbiomes empower Nicotiana tobacum complex traits dissection and prediction
Phyllosphere and rhizosphere microbiomes empower Nicotiana tobacum complex traits dissection and prediction
Du, Q.; Han, Y.; Liu, Z.; Si, H.; Ji, Y.; Liu, L.; Xiao, Z.; Cheng, L.; Yang, A.; Liu, D.; Zan, Y.
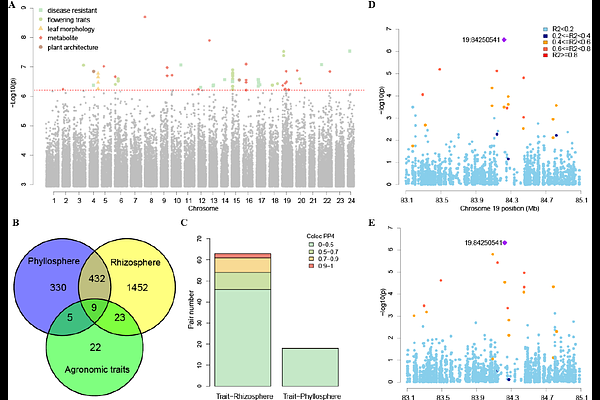
AbstractUnderstanding how plant-associated microbiomes interact with host genome variation to influence agronomic traits is essential for advancing microbiome-assisted crop improvement. In this study, we characterized the phyllosphere and rhizosphere microbiomes of 164 diverse Nicotiana tabacum accessions using 16S rRNA sequencing and integrated these data with host genomic variation and 22 agronomic traits. The two microbiomes exhibited distinct taxonomic structures, diversity patterns, and predicted metabolic functions. Microbiome genome-wide association studies identified extensive host genetic control over microbial abundance, including 49 shared genomic loci that explained nearly half of the heritable variation in both microbiomes. Microbiome-wide association studies revealed biologically meaningful associations between specific ASVs and agronomic traits. However, network analysis demonstrated that microbial sub-communities, rather than individual taxa, contributed substantially to phenotypic variation. Then, colocalization analysis further identified genetic variants jointly influencing microbial abundance and metabolite traits, highlighting potential host-microbe-trait causal links. Incorporating microbiome data into genomic selection models, we successfully improved prediction accuracy for several traits, especially plant architecture and flowering. Together, this work provides a comprehensive population-level framework linking host genetics, microbiome composition, and agronomic traits in tobacco, offering new insights for microbiome-informed breeding strategies.