A Multiscale Computational Architecture to Study Signaling Dynamics at Cell-Cell Interfaces
A Multiscale Computational Architecture to Study Signaling Dynamics at Cell-Cell Interfaces
Wu, Y.
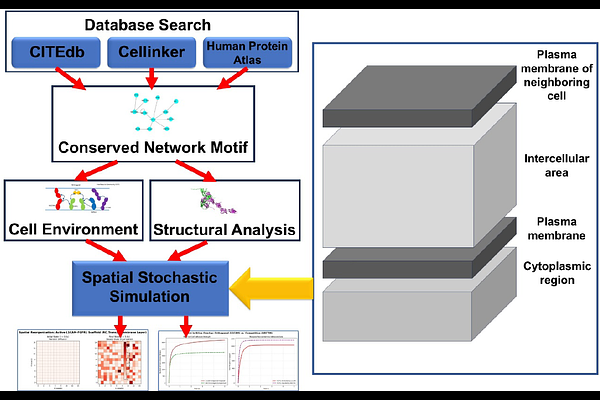
AbstractIntercellular communication is governed by the spatiotemporal dynamics of protein complexes at the cell-cell interface. However, conventional static interaction models fail to incorporate key physical constraints, such as steric hindrance, spatial compartmentalization, and dimensionality reduction that regulate complex assembly in vivo. To bridge the gap between static network topology and dynamic systems biology, we developed a multi-scale computational framework. We first identified a highly conserved, Fibroblast Growth Factor Receptor 1 (FGFR1)-centered cell adhesion and signaling motif by analyzing a diverse set of human cell-cell interfaces. We then constructed a multi-layer spatial stochastic simulator to recapitulate and interrogate the dynamic behavior of this network motif at cell-cell interfaces. Atomic-resolution structural models of the protein complexes within the motif were further generated using AlphaFold to define interaction rules for the stochastic simulations by categorizing binding interfaces. Our results show that the structural arrangement of cell-cell adhesion complexes controls how FGFR1 receptors cluster at the cell-cell interface, effectively dividing the membrane into distinct functional microdomains. Competition from decoy receptors further regulates this process by capturing receptors before they can participate in signaling. Even small changes in binding affinity can therefore alter receptor organization and disrupt normal signal transduction, which may contribute to human disease. By integrating macro-scale interactomics, atomic-level structural bioinformatics, and mesoscale stochastic modeling, this study reveals how structural interaction rules, combined with spatial constraints, shape the formation and function of intercellular signaling networks.