Selection mode governs the scaling of genetic load, diversity, and adaptation
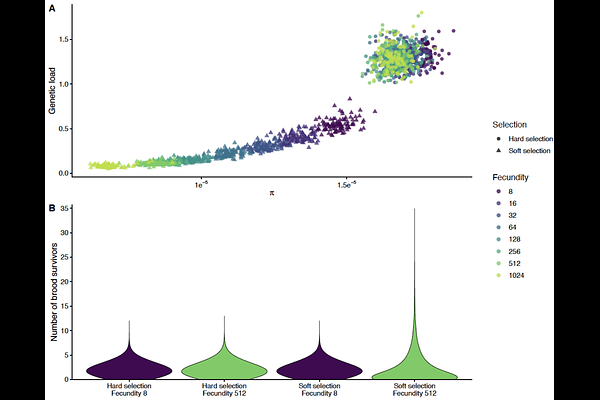
Selection mode governs the scaling of genetic load, diversity, and adaptation
Birley, T.; Oosterhout, C. v.
AbstractThe evolutionary consequences of selection depend on whether fitness is measured in absolute terms relative to a threshold (hard selection) or in relative terms among competing individuals (soft selection). Yet general predictions for how selection mode shapes the scaling of genome-wide diversity, genetic load, and adaptive change remain lacking. Using forward-time simulations, we show that hard and soft selection generate fundamentally different scaling relationships between population size, fecundity, nucleotide diversity, and genetic load. Under hard selection, increasing carrying capacity elevates both nucleotide diversity and load, consistent with the accumulation of mildly deleterious variation at larger effective sizes. Under soft selection, load rapidly approaches an asymptote as intensified within-cohort competition enhances purifying efficiency, while nucleotide diversity continues to increase, decoupling neutral diversity from mutational burden. High fecundity (r-strategy) strengthens this decoupling by promoting efficient purging but simultaneously generates extreme variance in reproductive success (sweepstakes), reducing effective population size (Ne) relative to census size (N) and limiting nucleotide diversity; despite large population numbers. Soft selection also enhances adaptive tracking under fluctuating phenotypic optima by avoiding the demographic costs associated with substitution load. Together, these results identify selection mode, alongside life history, as a key determinant of genome-wide load and diversity and provide a mechanistic explanation for why nucleotide diversity scales only weakly with census population size (Lewontins paradox).