Multicomplex Integrative Structural Modeling of a Human Histone Deacetylase Interactome
Multicomplex Integrative Structural Modeling of a Human Histone Deacetylase Interactome
Nde, J.; Majila, K.; Zimmermann, R.; Kempf, C.; Zhang, Y.; Cesare, J.; Thornton, J.; Workman, J.; Florens, L.; Viswanath, S.; Washburn, M.
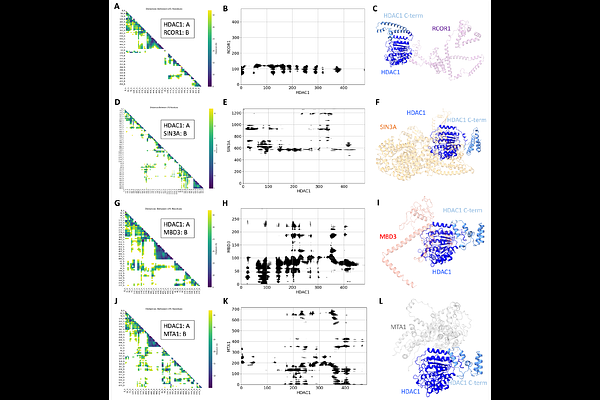
AbstractHistone Deacetylase (HDAC) 1 and 2 are key enzymatic components in multiple large chromatin remodeling complexes including NuRD, SIN3, and CoREST. In addition, both HDAC 1 and 2 contain a large intrinsically disordered region (IDR) within their C-terminal domain (CTD). How HDAC1/2 assemble into these complexes and the structure of the CTD IDR remains poorly understood. Here, we used HDAC1/2 to isolate their protein interaction networks from cells and used crosslinking mass spectrometry (XL-MS) coupled with the Integrative Modeling Platform to build structural models of the NuRD, SIN3A, and CoREST complexes. Next, we implemented an AlphaFold-enabled XL-MS constrained modeling approach to investigate how HDAC1 could assemble into these complexes. We show that the CTD IDR of HDAC1 folds into alpha helices in these complexes. Finally, we built a complete integrative structural model of a NuRD subcomplex including the abundant HDAC1:MBD3:MTA1:GATAD2B:RBBP4 subunits, which included 6 IDRs. The approaches used herein are broadly applicable for the study of protein complexes and protein interaction networks that can provide important insights into IDRs.