Kinetic Modeling of Target-Amplification-Free CRISPR-Cas-Based Autocatalysis Reactions
Kinetic Modeling of Target-Amplification-Free CRISPR-Cas-Based Autocatalysis Reactions
Wester, M.; Lim, J.; Van, A. B.; Koprowski, K.; Valera, E.; Bashir, R.
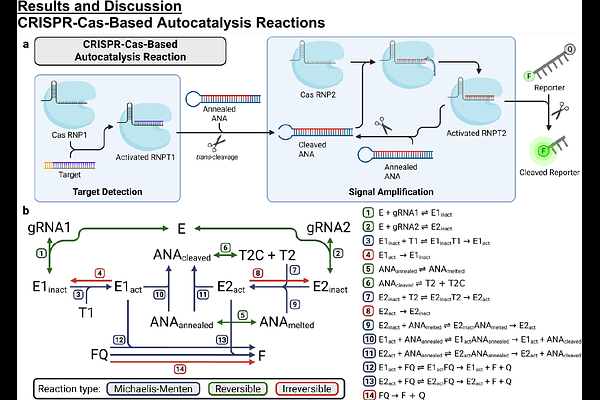
AbstractCRISPR-Cas-based diagnostics utilize the Cas enzyme's trans-cleavage activity to generate signal and have become popular platforms for sensitive nucleic acid detection. Recently, autocatalytic systems have been demonstrated to improve the time to response and sensitivity in some cases. However, mechanistic description of these assays is limited and optimization relies on simple trial-and-error. In this work, we present the first comprehensive kinetic model that integrates all major biochemical processes involved in these assays, including cleavage reactions, nucleic acid equilibrium kinetics, inhibition of trans-cleavage by single-stranded DNA, and degradation of single-stranded reaction components. We discuss the biochemical foundations and implementation of the ordinary differential equation model, which is built for adaptation to different reaction schemes. We use the full model to investigate the role of nucleic acid stability in assay performance for a typical nucleic acid design and show that our model demonstrates inhibition effects consistent with experimental data. We describe the reaction behavior, derive a simplified analytical model and compare its performance to the full analytical model. Finally, we demonstrate tools developed for rapid in silico optimization to guide the rational design of future target-amplification-free CRISPR-Cas-based autocatalysis assays.