Transcriptional profiling of extraocular motor neurons reveals sim1a as a candidate strabismus-related gene
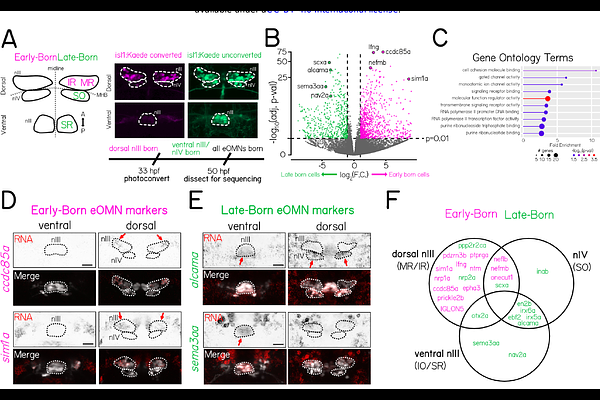
Transcriptional profiling of extraocular motor neurons reveals sim1a as a candidate strabismus-related gene
Gershowitz, E.; Hamling, K. R.; Rosti, B.; Gelnaw, H.; Xiang, G.; Quainoo, C.; Goldblatt, D.; Leary, P.; Schoppik, D.
AbstractStrabismus, or misalignment of the eyes, is a heritable disorder frequently associated with vision loss and decreased quality of life. Incomitant strabismus, where the degree of misalignment differs based on gaze angle, can arise from mutations in genes that regulate the development of extraocular motor neurons. To date, few such genes have been identified. The extraocular motor system is highly conserved across vertebrates, suggesting a comparative transcriptomic discovery approach would be fruitful. Using bulk and single-cell sequencing in a small accessible vertebrate, the larval zebrafish, we identified genes expressed in subpopulations of extraocular motor neurons in cranial nuclei nIII/nIV. We next assessed extraocular motor neuron number and vestibulo-ocular reflex performance after CRISPR/Cas9-mediated mutagenesis of three genes with suggestive expression patterns: sim1a, nav2a, onecut1, and one known to disrupt nIII/nIV motor neuron specification: phox2a. Loss of sim1a impaired the vestibulo-ocular reflex without change to nIII/nIV motor neuron number. Our data suggest that constitutive disruptions to sim1a can impair nIII/nIV-dependent eye movements. More broadly, our work illuminates considerable transcriptomic diversity among extraocular motor neuron subpopulations, and establishes a pipeline to identify genes relevant to ocular motor disease etiology.