A Deep Learning Framework for Predicting Gut Microbe-Host Receptor Interactions
A Deep Learning Framework for Predicting Gut Microbe-Host Receptor Interactions
li, H.; Zhao, R.; Zhu, C.; Jiang, R.; Chen, T.; li, X.; Yang, Y.
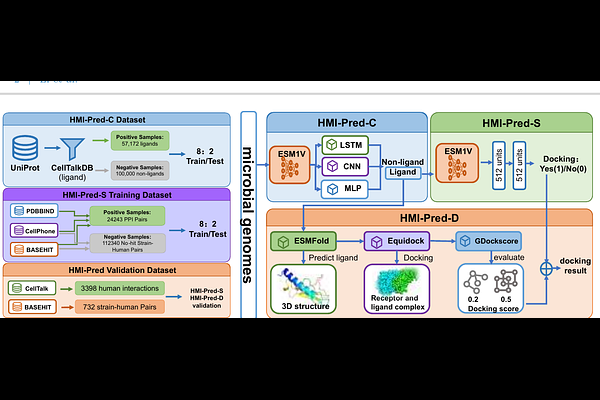
AbstractGut microbiota regulates host health through complex protein-protein interactions. However, deciphering this specific interactions between microbiota and human receptors remains a significant challenge due to the lack of specialized computational tools. Leveraging the hypothesis of cell communication and relevant data, HMI-Pred initially builds an ensemble classifier to screen for potential ligand sequences within microbial genomes. It then jointly evaluates sequence semantics and molecular docking to predict potential microbe-host receptor interactions. HMI-Pred achieved robust performance with F1-scores of 0.901 for microbial ligand identification and 0.883 for interaction prediction. Application to 332,381 microbial proteins revealed distinct interaction patterns: histone deacetylases (HDACs) served as broad-spectrum targets (mean score >0.80), while G protein-coupled receptors (GPCRs) exhibited high specificity (scores 0.42-0.61). Furthermore, literature mining validated over 47% of the functional predictions, and specific immunomodulatory interactions were confirmed in Akkermansia muciniphila. HMI-Pred provides a valuable computational tool for decoding host-microbe signaling networks and facilitating the discovery of microbiome-based therapeutic targets.