From genes to collective modes: biological constraints shape metabolic evolution
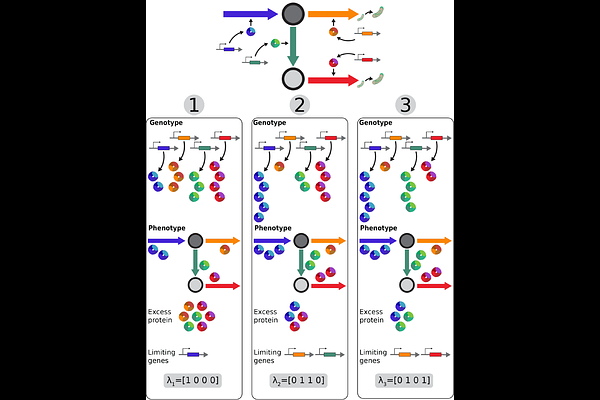
From genes to collective modes: biological constraints shape metabolic evolution
Brown, A.; Datta, S.; Mehta, P.; Cleary, B.
AbstractPopulation genetics has been successful in explaining how drift, selection, and mutation shape allele frequencies. However, we still lack an understanding of how biological and evolutionary constraints arising from genotype-phenotype maps shape evolutionary dynamics. To explore this question, we developed a new framework for simulating the evolution of an inherently polygenic and epistatic trait - metabolism - combining population genetics with flux balance analysis. We found that the evolution of metabolic networks exhibits surprisingly simple, reproducible dynamics. We identify evolutionary collective modes (EvCMs) - linear combinations of genes with high, constant selective pressure - as organizers of the long-term dynamics and the origin of this simplicity. EvCMs arise through the interaction of physical constraints, evolvability, and the requirements for growth. We developed a theoretical framework to predict EvCMs from large-scale metabolic models and found quantitative agreement between theory and simulation of E. coli central metabolism. Inspired by these results, we re-analyzed mutational data from Lenski's long-term evolution experiment and found compelling evidence for the existence of EvCMs. This work suggests that biological constraints encoded in genotype-phenotype maps play an important role in shaping evolutionary dynamics, offering a new perspective on complex trait evolution - one that shifts focus from individual genes to collective modes.