Assembling a fully-dated complete tree of life
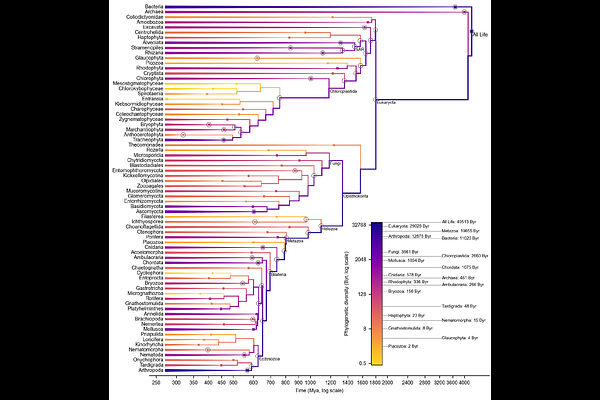
Assembling a fully-dated complete tree of life
Duke, J. D.; Guo, J.; Forest, F.; Gumbs, R.; McTavish, E. J.; Rosindell, J.
AbstractTime-scaled phylogenetic trees summarising evolutionary relationships are fundamental to many analyses in biology, from diversification rate estimation to conservation prioritisation. The most comprehensive available summary of these relationships, the Open Tree of Life, synthesises information from over two thousand studies into a supertree covering the full range of global biodiversity, but its use in downstream analyses is limited by the lack of divergence times. Previous work has mapped dates from Open Tree's database of trees to certain nodes in the supertree, but for the majority of nodes no date is available. While algorithms exist to interpolate missing dates in a tree, we found that their time and memory requirements scaled quadratically with the number of nodes, which made it computationally infeasible to run them on the entire tree. In this work, we describe novel date interpolation algorithms that scale linearly with the number of nodes. These enabled us to produce a distribution of fully-dated trees containing 2.3 million extant described species, greatly expanding the scope of feasible phylogenetic analyses. We illustrate the utility of these trees by computing the most robust estimate yet of the phylogenetic diversity of the complete tree of life, incorporating both topological and temporal uncertainty.